An Online Digital Critical Apparatus
The online Apparatus Explorer is a web app implementation of my standalone desktop version.
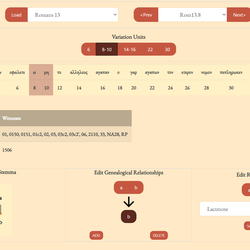
[NOTE: Apparatus Explorer has been entirely redesigned and renamed to Apatosaurus. The new version includes a comprehensive user interface for the CBGM. For more on the new version, see its About page.]
A Little Background on the Apparatus Explorer
One of the tools I use every day while I'm writing up my doctoral thesis is a little desktop app (Apparatus Explorer) that I developed for easily visualizing and editing an XML TEI encoded digital critical apparatus. TEI is human readable on its own, indeed, TEI encoded documents are commonly produced by humanities scholars whose specialty is far from software development. On the other hand, non-styled TEI is not really meant to be published 'as is,' but its value comes from carrying additional data along with the plain transcription (corrections, reading types, etc.). The Apparatus Explorer is one way to take the text and extra encoded information from a collation file and display it back to a human editor as an intuitive and useful digital critical apparatus.
What the Apparatus Explorer Does
In this online version of the Apparatus Explorer, a (logged in) user can upload a collation file. The file will then go through some processing in the server:
- The file is cleaned and reformatted using Joey McCollum's helpful script.
- Since the collation file will usually be the output of ITSEE's Collation Editor (as all of mine are), I've written a module that renames the identifiers for readings which have been assigned as subreadings of a parent reading. By default, the Collation Editor assigns all subreadings of a parent reading to the same name (e.g., all subreadings of reading 'a' are identified as 'ar').
From here the files are ready to be viewed in the Apparatus Explorer. If any local stemmata have been encoded, these are displayed as graphs. In fact, one of the main goals for this project was to easily encode (i.e. by clicking buttons rather than hand encoding) local stemmata for each variant in preparation for using the collation file as input for the open-cbgm. Additionally, the Apparatus Explorer enables simple addition of reading types (e.g. lacunose, orthographic, etc.). All of this is built into the desktop app.
I am very pleased to offer the same functionality as a web app. A web app has several advantages over the desktop application:
- A web app provides me (and others!) with a way to publish critical apparatuses in a uniquely digital medium. The initial inspiration for this project was my desire to deliver my research data to my doctoral supervisor in a more useful format.
- Web apps are cross platform; the desktop app only worked well on Windows, but I have designed the web app to work in both desktop and mobile browsers. (I haven't tested it on Safari, so I'd would love to hear from others if it looks and works fine).
Check out the Demo!
I'm sure you will understand why only logged in users may upload their own collation files. However, I have one of my own collation files uploaded and ready to be parsed and viewed.
Demo Tutorial
[NOTE: I no longer host a demo version, but instead, you can see published examples on the Apparatus Explorer's new home: https://apparatusexplorer.com/]
1. select an available collation and click 'Load.' At the time of writing this post I've only added one to the demo.
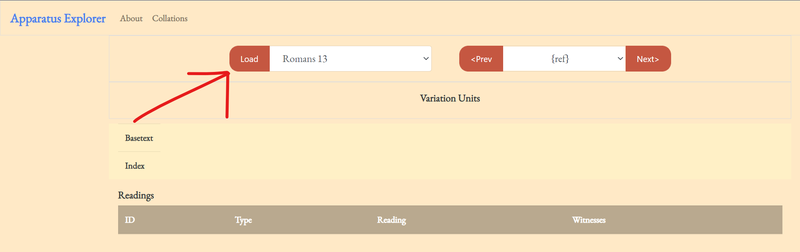
If the collation file contains multiple verses (I work a chapter at a time), any verse can be selected from the dropdown menu or adjacent verses can be selected by clicking Prev and Next.
Upon loading, the variation units, basetext, and indices should be generated. Think of this as similar to the ECM's layout. Each word in the basetext is given an index number, so every variation unit can be identified as a range of these index numbers. The basetext is reconstructed from the TEI file by piecing together all the readings tagged as belonging to the basetext. In the case of the example file, the NA28 was used as the basetext.
2. click on one of the variation units.
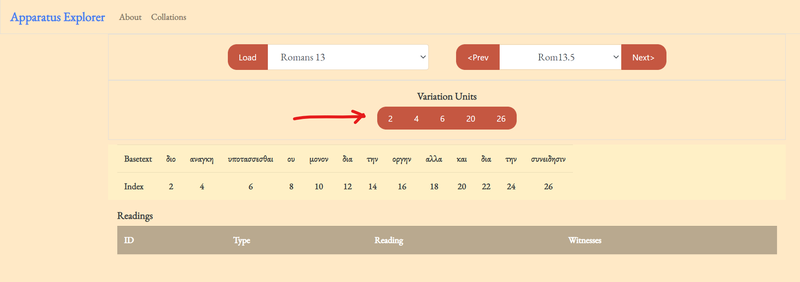
Upon clicking on a variation unit, the rest of the explorer will generate. Notice that the selected unit is highlighted in the basetext.
3. Add a local stemma information by clicking on a reading to designate as the 'from' node.
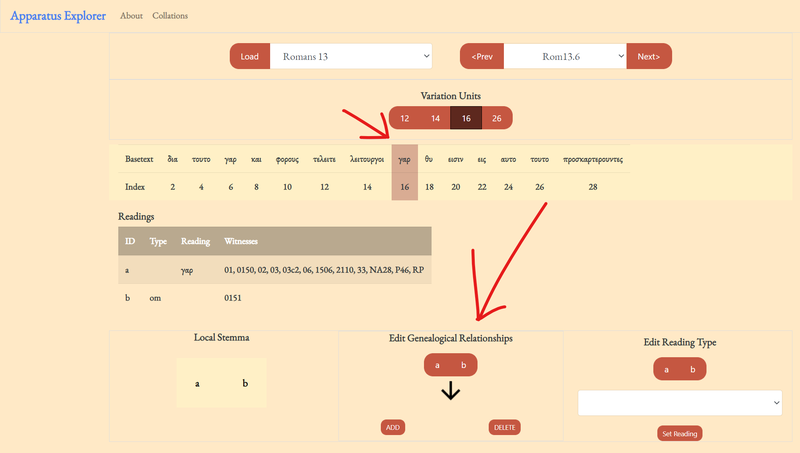
After clicking on a 'from' or starting node, the available 'to' nodes will appear. Make your selections and then click 'ADD.' You should see the graph update; when the verse is saved (a feature not available in the demo), the original collation file will be updated with the edits. These choices become powerful in combination with the open-cbgm.
4. Change or add a reading type.
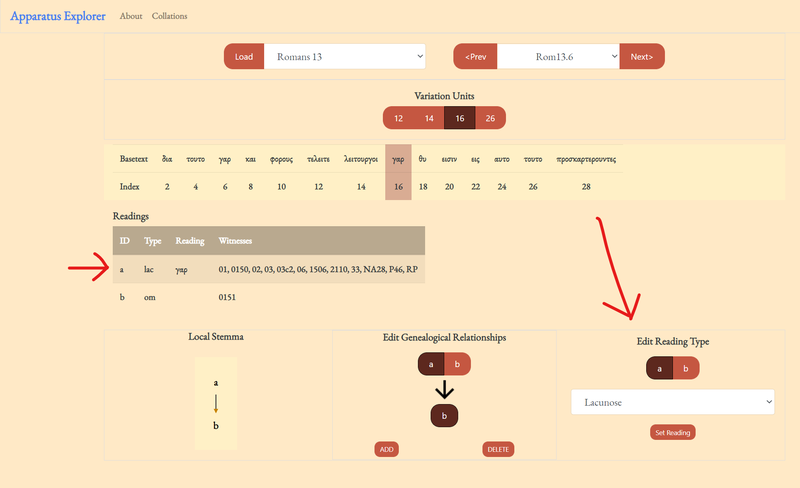
Finally, update a reading by first selecting the reading identifier, selecting a reading type, and then clicking 'ADD.' Most readings do not need a type encoded, but they are helpful for several things. First, adding a type helps to clearly distinguish between lacunose and omitted text. Second, the open-cbgm can be configured to count different reading types in different ways. For example, one may want to exclude minor spelling differences before calculating pregenealogical coherence. If that is the case, then readings labeled "orthographic" will be considered to agree with their parent reading.
Future Plans
- Export the apparatus as a Microsoft Word docx file and xlsx file (this is already working; I just need to incorporate that code into the backend).
- Lighten up the front end. There is a lot more JavaScript to load than is required. I picked up some software development skills only a year ago as a hobby during the first pandemic lockdown. I spent the year learning Python (the backend of the apparatus explorer is Django) and just enough HTML and CSS, but only a wee bit of JavaScript. If I have time to pick it up better, hopefully I can start dropping the various libraries required to run the Apparatus Explorer as it is. [This has been resolved in the new version at https://apparatusexplorer.com/].
- I've thought about adding the ability to edit text readings and witnesses in the collation file. The reason I've avoided this, though, is that if there is an error, it probably goes back to a transcription. I think it is better practice to go and fix the transcription and recollate that verse. This ensures accurate documents from beginning to end.
Feedback is appreciated!
This is a niche tool. It is limited to those who have their own TEI collation files, encoded by hand or from the Collation Editor. But if this tool finds its way to another person doing similar research, please let me know what works and what does not.
I would love to see other critical apparatuses displayed in this way.